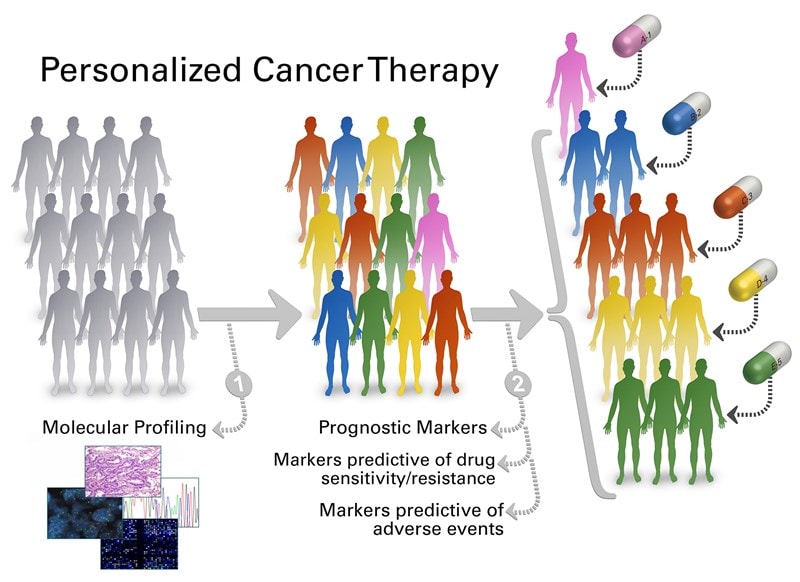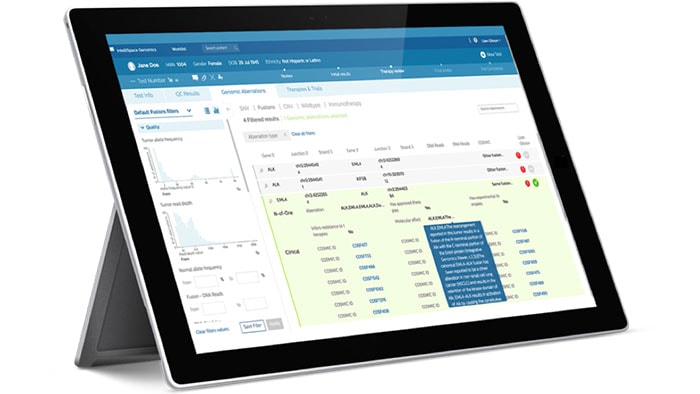
To keep pace with the rapid developments in the field of genomics, you need solutions that incorporate the evolving genomic data, can articulate your health system workflows in pathology and oncology, and integrate deeply with your current technology. Whether it be executing user-defined genomic workflows, interpreting clinical reports, biomarker-based clinical trial matching, or understanding your cancer population– our solutions can help.
Why Philips Genomics
Genomics analysis
Offers a controlled bioinformatics environment for running pipelines. It offers traceability, reproducibility, QC, storage, and a versatile pipeline infrastructure that supports both customized and dockerized pipeline, for cancer genomic data. The data may include .fastq, .bcl formats, as well as .vcf , .json and .xml.
Automated end-to-end workflow
Automating pathologists’ workflow from loading sequencing output files to customized report generation, including therapy options and trial matching with expert curated clinical knowledgebase.
Secure, sustainable, and scalable infrastructure
Dependable mechanism for automatic, secure, quality assured acquisition of clinical and genomic data. Distributed software system with microservices.
Comprehensive variant review and filter
Automated and streamlined variant calling and prioritization using leading variant annotation databases and clinical evidence.
Vendor agnostic integration
Automated clinical data inbound/outbound integration per specification with our IntelliBridge Enterpise integration engine from/to EHR or other clinical IT systems.
Multiple knowledgebases
Comprehensive clinical knowledgebases of most up-to-date literature, therapies and clinical trials from Qiagen N-of-One, Jackson Laboratory, and MD Anderson.
Bridging pathology to oncology

Vendor agnostic

Multiple knowledgebases
Connecting in-house
Delivering an automated end-to-end workflow
Genomics in your tumor board experience

Our molecular tumor board provides your oncologists a unified view of relevant therapies and clinical trials in the context of each patient’s unique genomic profile, which can help your clinicians make evidence-based decisions on patient treatment.
For more information on our Genomics Workspace
Philips Genomics solutions in the news

Journal of Precision Medicine
Philips vision is to become a leader in innovating precision medicine solutions that connect areas such as pathology, genomics, and molecular phenotyping molecular for personalized therapy decision making. To learn more about this platform, we connected with Louis Culot, General Manager, Oncology Informatics and Genomics, and Dr. Qi Wei, Genomic Subject Matter Expert.

Journal of Clinical Pathways
Journal of Clinical Pathways spoke with Kenna Shaw, PhD, cancer genomics laboratory, MD Anderson, and Louis Culot, general manager, genomics and oncology informatics, Philips, to better understand the PODS system and how this collaboration will improve patient care worldwide.

Genomics: Decoding the future
Jeroen Tas, Chief Innovation & Strategy Officer of Philips, shares his insights on the current and future role of genomics in medical practices and treatment selection.
“We are right in the middle of a genomic revolution.”
Our Oncology informatics portfolio
Genomics Workspace current product name is Pathology Genomics Workspace and Oncology Genomics Workspace. The name of the product will change to Genomics Workspace with the release of ISPM V5.3, subject to regulatory approval.
Oncology Pathways is also known as the Oncology Pathways Navigator. The Oncology Pathways name will be applicable as of ISPM V5.3.